Pyrrole-6-6YV4
IC50 6.3 +/- 0.5 microM Notum with OPTS substrate
General
Type : Cyclopropyl,Pyrrole_Pyrrolidine-derivative
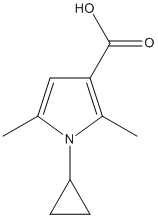
Chemical_Nomenclature : 1-cyclopropyl-2,5-dimethylpyrrole-3-carboxylic acid
Canonical SMILES : CC1=CC(=C(N1C2CC2)C)C(=O)O
InChI : InChI=1S\/C10H13NO2\/c1-6-5-9(10(12)13)7(2)11(6)8-3-4-8\/h5,8H,3-4H2,1-2H3,(H,12,13)
InChIKey : VOOZJPFNBFNPEK-UHFFFAOYSA-N
Other name(s) : 1-cyclopropyl-2,5-dimethyl-1H-pyrrole-3-carboxylic acid,1-cyclopropyl-2,5-dimethyl-pyrrole-3-carboxylic Acid,MLS000773859,PQK,Pyrrole-3-carboxylic acid fragment 686,SCHEMBL1714277,CHEMBL1577069,CTK1D5460,ZINC159030,DB-015731
MW : 179.22
Formula : C10H13NO2
CAS_number : 423768-58-5
PubChem : 2776579
UniChem : VOOZJPFNBFNPEK-UHFFFAOYSA-N
IUPHAR :
Wikipedia :
Target
Families : Pyrrole-6-6YV4 ligand of proteins in family: Pectinacetylesterase-Notum
Stucture : 6YV4 Structure of the human Wnt deacylase Notum in complex with a pyrrole-3-carboxylic acid fragment 454
Protein : human-NOTUM
References (1)
| Title : Screening of a Custom-Designed Acid Fragment Library Identifies 1-Phenylpyrroles and 1-Phenylpyrrolidines as Inhibitors of Notum Carboxylesterase Activity - Mahy_2020_J.Med.Chem_63_9464 |
| Author(s) : Mahy W , Patel M , Steadman D , Woodward HL , Atkinson BN , Svensson F , Willis NJ , Flint A , Papatheodorou D , Zhao Y , Vecchia L , Ruza RR , Hillier J , Frew S , Monaghan A , Costa A , Bictash M , Walter MW , Jones EY , Fish PV |
| Ref : Journal of Medicinal Chemistry , 63 :9464 , 2020 |
| Abstract : Mahy_2020_J.Med.Chem_63_9464 |
| ESTHER : Mahy_2020_J.Med.Chem_63_9464 |
| PubMedSearch : Mahy_2020_J.Med.Chem_63_9464 |
| PubMedID: 32787107 |
| Gene_locus related to this paper: human-NOTUM |