4HS9
Name : Methanol tolerant mutant of the Proteus mirabilis lipase
Revelation date : 22-May-2013
Family : Bacterial_lip_FamI.1
Gene_locus : promi-c2lfd0
PDB file : ESTHER: header of PDB entry RCSB: Full entry
Comment
Korman, T.P., Sahachartsiri, B., Charbonneau, D.M., Huang, G.L., Beauregard, M., Bowie, J.U. 13 mutations G181C\/S238C\/K208N\/L64I\/A70T\/F225L\/Q277L\/G202E\/G266S\/D270N\/N17S\/I255F\/R33T
References (1)
| Title : Dieselzymes: development of a stable and methanol tolerant lipase for biodiesel production by directed evolution. - Korman_2013_Biotechnol.Biofuels_6_70 |
| Author(s) : Korman TP , Sahachartsiri B , Charbonneau DM , Huang GL , Beauregard M , Bowie JU |
| Ref : Biotechnol Biofuels , 6 :70 , 2013 |
| Abstract : |
| PubMedSearch : Korman_2013_Biotechnol.Biofuels_6_70 |
| PubMedID: 23648063 |
| Gene_locus related to this paper: promi-c2lfd0 |
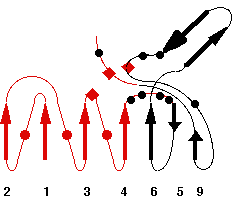
Representative scheme of Prolylcarboxypeptidase structure and an image from PDBsum server

Databases
PDB-Sum : 4HS9 Previously Class, Architecture, Topology and Homologous superfamily - PDB-Sum server
FSSP : 4HS9 Fold classification based on Structure-Structure alignment of Proteins - FSSP server
SCOP : 4HS9 Structural Classification Of Protein - SCOP server
PROTEOPEDIA : 4HS9 3D, interactive encyclopedia of proteins - PROTEOPEDIA server