2ZJ7
Name : Crystal structure of D157A mutant of Pseudomonas sp. MIS38 lipase
Revelation date : 02-Dec-2008
Family : Bacterial_lip_FamI.3
Gene_locus : psesp-Q9RBY1
PDB file : ESTHER: header of PDB entry RCSB: Full entry
Comment
Kuwahara, K, Angkawidjaja, C, Matsumura, H, Koga, Y, Takano, K, Kanaya, S
Ligand : Ligand in structure: Ligplot
References (1)
| Title : Importance of the Ca2+-binding sites in the N-catalytic domain of a family I.3 lipase for activity and stability - Kuwahara_2008_Protein.Eng.Des.Sel_21_737 |
| Author(s) : Kuwahara K , Angkawidjaja C , Matsumura H , Koga Y , Takano K , Kanaya S |
| Ref : Protein Engineering Des Sel , 21 :737 , 2008 |
| Abstract : |
| PubMedSearch : Kuwahara_2008_Protein.Eng.Des.Sel_21_737 |
| PubMedID: 18987131 |
| Gene_locus related to this paper: psesp-Q9RBY1 |
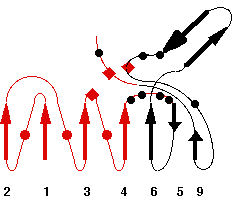
Representative scheme of Prolylcarboxypeptidase structure and an image from PDBsum server

Databases
PDB-Sum : 2ZJ7 Previously Class, Architecture, Topology and Homologous superfamily - PDB-Sum server
FSSP : 2ZJ7 Fold classification based on Structure-Structure alignment of Proteins - FSSP server
SCOP : 2ZJ7 Structural Classification Of Protein - SCOP server
PROTEOPEDIA : 2ZJ7 3D, interactive encyclopedia of proteins - PROTEOPEDIA server