4GXN
Name : Diethylphosphonate Inhibited Structure of the Proteus mirabilis Lipase
Revelation date : 06-Feb-2013
Family : Bacterial_lip_FamI.1
Gene_locus : promi-c2lfd0
PDB file : ESTHER: header of PDB entry RCSB: Full entry
Comment
Korman, T.P., Bowie, J.U.
Ligand : Inhibitor Paraoxon, Diethyl-hydrogen-phosphate Ligand in structure: Ligplot
References (1)
| Title : Crystal Structure of Proteus mirabilis Lipase, a Novel Lipase from the Proteus\/Psychrophilic Subfamily of Lipase Family I.1 - Korman_2012_PLoS.One_7_e52890 |
| Author(s) : Korman TP , Bowie JU |
| Ref : PLoS ONE , 7 :e52890 , 2012 |
| Abstract : |
| PubMedSearch : Korman_2012_PLoS.One_7_e52890 |
| PubMedID: 23300806 |
| Gene_locus related to this paper: promi-c2lfd0 |
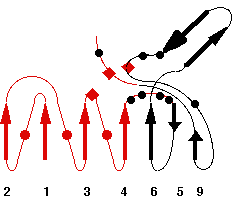
Representative scheme of Prolylcarboxypeptidase structure and an image from PDBsum server

Databases
PDB-Sum : 4GXN Previously Class, Architecture, Topology and Homologous superfamily - PDB-Sum server
FSSP : 4GXN Fold classification based on Structure-Structure alignment of Proteins - FSSP server
SCOP : 4GXN Structural Classification Of Protein - SCOP server
PROTEOPEDIA : 4GXN 3D, interactive encyclopedia of proteins - PROTEOPEDIA server