4UBJ
Name : Kinetic crystallography of alphaE7-carboxylesterse from Lucilia cuprina.Absorbed X-ray dose 5.55 MGy at 100K
Revelation date : 19-Aug-2015
Family : Carb_B_Arthropoda
Gene_locus : luccu-E3aest7
PDB file : ESTHER: header of PDB entry RCSB: Full entry
Comment
Jackson, C.J., Carr, P.D., Weik, M., Huber, T., Meirelles, T., Correy, G.
Ligand :
References (1)
| Title : Mapping the Accessible Conformational Landscape of an Insect Carboxylesterase Using Conformational Ensemble Analysis and Kinetic Crystallography - Correy_2016_Structure_24_977 |
| Author(s) : Correy GJ , Carr PD , Meirelles T , Mabbitt PD , Fraser NJ , Weik M , Jackson CJ |
| Ref : Structure , 24 :977 , 2016 |
| Abstract : Correy_2016_Structure_24_977 |
| ESTHER : Correy_2016_Structure_24_977 |
| PubMedSearch : Correy_2016_Structure_24_977 |
| PubMedID: 27210287 |
| Gene_locus related to this paper: luccu-E3aest7 |
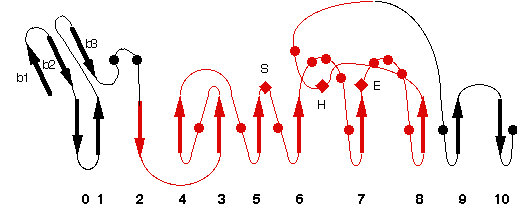
Representative scheme of Prolylcarboxypeptidase structure and an image from PDBsum server