Paranitrophenyl-decanoate
General
Type : Water-insoluble medium and long chain ester || pNP || Chromogen || Decanoate
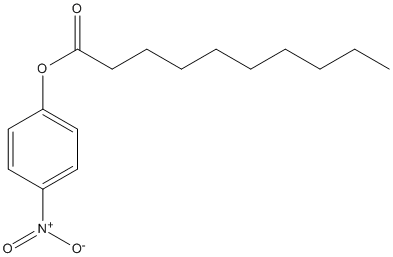
Chemical_Nomenclature : (4-nitrophenyl) decanoate
Canonical SMILES : CCCCCCCCCC(=O)OC1=CC=C(C=C1)[N+](=O)[O-]
InChI : InChI=1S\/C16H23NO4\/c1-2-3-4-5-6-7-8-9-16(18)21-15-12-10-14(11-13-15)17(19)20\/h10-13H,2-9H2,1H3
InChIKey : RRNKCSIEGPOIKI-UHFFFAOYSA-N
Other name(s) : pNP-C10, 4-Nitrophenyl decanoate, Decanoic acid 4-nitrophenyl ester, (4-nitrophenyl) Decanoate, p-NP caprate, pNP-caprate, pNP-decanoate, p-nitrophenyl decanoate, p-NP decanoate, P-nitrophenyl caprate, 4-Nitrophenyl caprate, p-NPD
References (5)
| Title : Directed evolution of new and improved enzyme functions using an evolutionary intermediate and multidirectional search - Porter_2015_ACS.Chem.Biol_10_611 |
| Author(s) : Porter JL , Boon PL , Murray TP , Huber T , Collyer CA , Ollis DL |
| Ref : ACS Chemical Biology , 10 :611 , 2015 |
| Abstract : Porter_2015_ACS.Chem.Biol_10_611 |
| ESTHER : Porter_2015_ACS.Chem.Biol_10_611 |
| PubMedSearch : Porter_2015_ACS.Chem.Biol_10_611 |
| PubMedID: 25419863 |
| Gene_locus related to this paper: psepu-clcd1 |
| Title : The metagenome-derived enzymes LipS and LipT increase the diversity of known lipases - Chow_2012_PLoS.One_7_e47665 |
| Author(s) : Chow J , Kovacic F , Dall Antonia Y , Krauss U , Fersini F , Schmeisser C , Lauinger B , Bongen P , Pietruszka J , Schmidt M , Menyes I , Bornscheuer UT , Eckstein M , Thum O , Liese A , Mueller-Dieckmann J , Jaeger KE , Streit WR |
| Ref : PLoS ONE , 7 :e47665 , 2012 |
| Abstract : Chow_2012_PLoS.One_7_e47665 |
| ESTHER : Chow_2012_PLoS.One_7_e47665 |
| PubMedSearch : Chow_2012_PLoS.One_7_e47665 |
| PubMedID: 23112831 |
| Gene_locus related to this paper: symth-q67mr3 , 9bact-k7qe48 |
| Title : Optimization of a thermostable lipase from Bacillus stearothermophilus P1: overexpression, purification, and characterization - Sinchaikul_2001_Protein.Expr.Purif_22_388 |
| Author(s) : Sinchaikul S , Sookkheo B , Phutrakul S , Pan FM , Chen ST |
| Ref : Protein Expr Purif , 22 :388 , 2001 |
| Abstract : Sinchaikul_2001_Protein.Expr.Purif_22_388 |
| ESTHER : Sinchaikul_2001_Protein.Expr.Purif_22_388 |
| PubMedSearch : Sinchaikul_2001_Protein.Expr.Purif_22_388 |
| PubMedID: 11483000 |
| Gene_locus related to this paper: geost-lipas |
| Title : A novel extracellular esterase from Bacillus subtilis and its conversion to a monoacylglycerol hydrolase - Eggert_2000_Eur.J.Biochem_267_6459 |
| Author(s) : Eggert T , Pencreac'h G , Douchet I , Verger R , Jaeger KE |
| Ref : European Journal of Biochemistry , 267 :6459 , 2000 |
| Abstract : Eggert_2000_Eur.J.Biochem_267_6459 |
| ESTHER : Eggert_2000_Eur.J.Biochem_267_6459 |
| PubMedSearch : Eggert_2000_Eur.J.Biochem_267_6459 |
| PubMedID: 11029590 |
| Gene_locus related to this paper: bacsu-LIPB |
| Title : Purification and partial characterization of a novel thermophilic carboxylesterase with high mesophilic specific activity - Wood_1995_Enzyme.Microb.Technol_17_816 |
| Author(s) : Wood AN , Fernandez-Lafuente R , Cowan DA |
| Ref : Enzyme Microb Technol , 17 :816 , 1995 |
| Abstract : Wood_1995_Enzyme.Microb.Technol_17_816 |
| ESTHER : Wood_1995_Enzyme.Microb.Technol_17_816 |
| PubMedSearch : Wood_1995_Enzyme.Microb.Technol_17_816 |
| PubMedID: 7576531 |
| Gene_locus related to this paper: geost-est50 |